Home » Student Resources » Online Chemistry Textbooks » CH450 and CH451: Biochemistry - Defining Life at the Molecular Level » Chapter 4: DNA, RNA, and the Human Genome
MenuCH450 and CH451: Biochemistry - Defining Life at the Molecular Level
Chapter 4: DNA, RNA, and the Human Genome
4.1 The Structure of DNA and RNA
4.2 Chromosomes and Packaging
4.3 Sequencing the Human Genome
4.4 References
4.1 The Structure of DNA and RNA
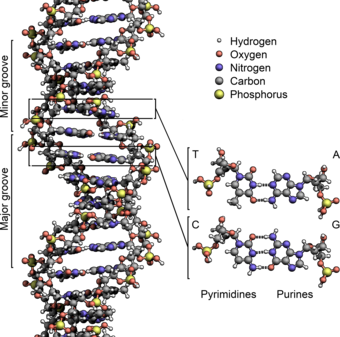
Alongside proteins, lipids and complex carbohydrates (polysaccharides), nucleic acids are one of the four major types of macromolecules that are essential for all known forms of life. The nucleic acids consists of two major macromolecules, Deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) that carry the genetic instructions for the development, functioning, growth and reproduction of all known organisms and viruses. The DNA macromolecule (Figure 4.1) is composed of two polynucleotide chains that coil around each other to form a double helix. The RNA macromolecule usually exists as a single polynucleotide chain that is much shorter than the comparative DNA molecule.

Figure 4.1: The structure of the DNA double helix. The atoms in the structure are color-coded by element and the detailed structures of two base pairs are shown in the bottom right.
Image on the left by: Zephyris
DNA Animation on the right by:
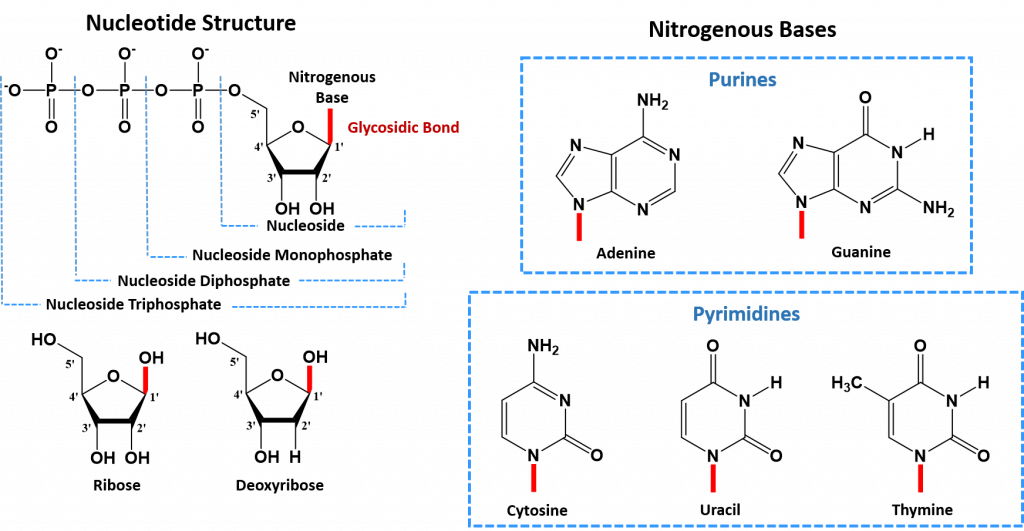
The core structure of a nucleic acid monomer is the nucleoside, which consists of a sugar residue + a nitrogenous base that is attached to the sugar residue at the 1′ position (Figure 4.2). The sugar utilized for RNA monomers is ribose, whereas DNA monomers utilize deoxyribose that has lost the hydroxyl functional group at the 2′ position of ribose. For the DNA molecule, there are four nitrogenous bases that are incorporated into the standard DNA structure. These include the Purines: Adenine (A) and Guanine (G), and the Pyrimidines: Cytosine (C) and Thymine (T). RNA uses the same nitrogenous bases as DNA, except for Thymine. Thymine is replaced with Uracil (U) in the RNA structure.
When one or more phosphate groups are attached to a nucleoside at the 5′ position of the sugar residue, it is called a nucleotide. Nucleotides come in three flavors depending how many phosphates are included: the incorporation of one phosphate forms a nucleoside monophosphate, the incorporation of two phosphates forms a nucleoside diphosphate, and the incorporation of three phosphates forms a nucleoside triphosphate (Figure 4.2).
Figure 4.2 The Monomer Building Blocks of Nucleic Acids. The site of the nitrogenous base attachment to the sugar residue (glycosidic bond) is shown in red.
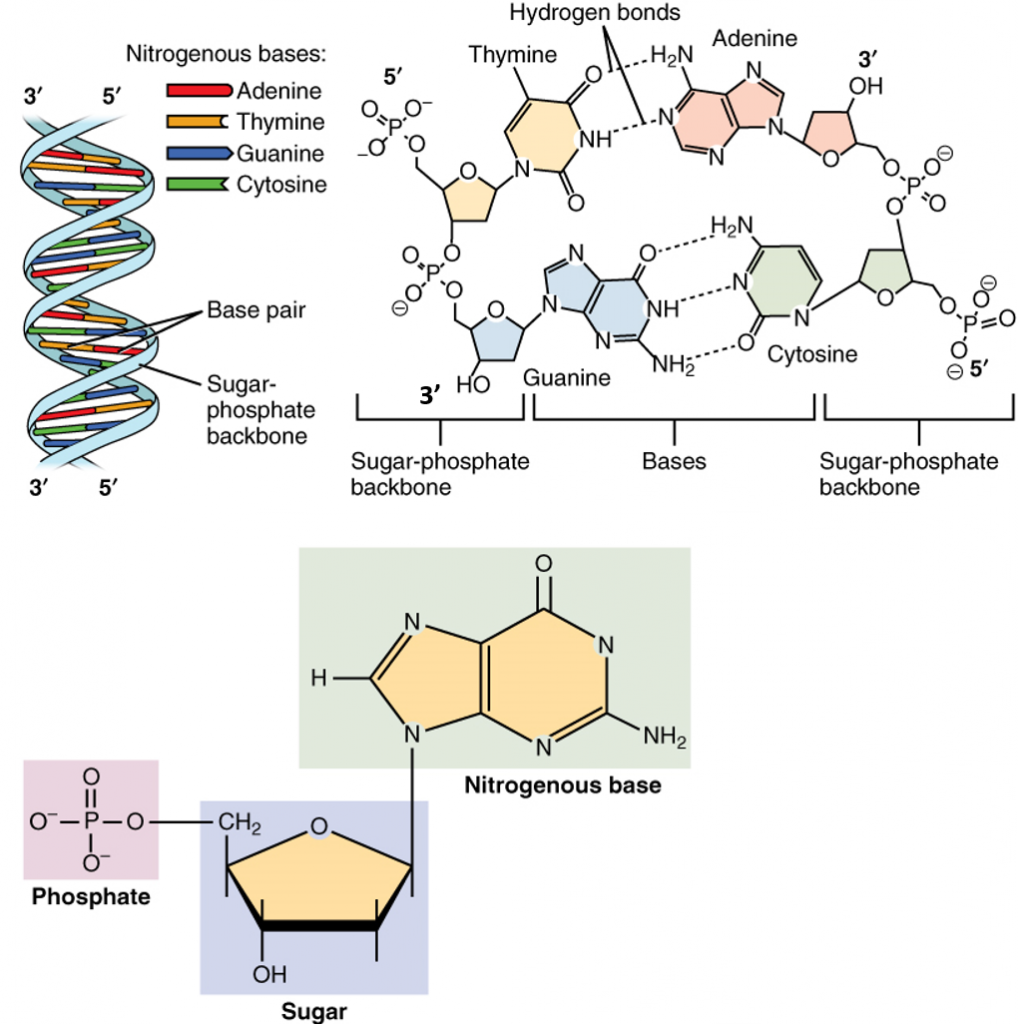
The double helix formed during DNA synthesis has several key physical properties (Figure 4.3). DNA is assembled such that nucleoside monophosphates are incorporated into the growing DNA chains. Unlike the protein α-helix, where the R-groups of the amino acids are positioned to the outside of the helix, in the DNA double helix, the nitrogenous bases are positioned inward and face each other. The backbone of the DNA is made up of repeating sugar-phosphate-sugar-phosphate residues. Bases fit in the double helical model if pyrimidine on one strand is always paired with purine on the other. From Chargaff’s rules, the two strands will pair A with T and G with C. This pairs a keto base with an amino base, a purine with a pyrimidine. Two H‑bonds can form between A and T, and three can form between G and C. This third H-bond in the G:C base pair is between the additional exocyclic amino group on G and the C2 keto group on C. The pyrimidine C2 keto group is not involved in hydrogen bonding in the A:T base pair.
Furthermore, the orientation of the sugar molecule within the strand determines the directionality of the strands. The phosphate group that makes up part of the nucleotide monomer is always attached to the 5′ position of the deoxyribose sugar residue. The free end that can accept a new incoming nucleotide is the 3′ hydroxyl position of the deoxyribose sugar. Thus, DNA is directional and is always synthesized in the 5′ to 3′ direction. Interestingly, the two strands of the DNA double helix lie in opposite directions or have a head to tail orientation.
Figure 4.3 Structure of DNA: Lower diagram shows the arrangement of the nucleoside monophosphate within the structure of nucleic acids. In the upper right four nucleotides form two base-pairs: thymine and adenine (connected by double hydrogen bonds) and guanine and cytosine (connected by triple hydrogen bonds). The individual nucleotide monomers are chain-joined at their sugar and phosphate molecules, forming two ‘backbones’ (a double helix) of a nucleic acid, shown at upper left.
Image modified from: Openstax
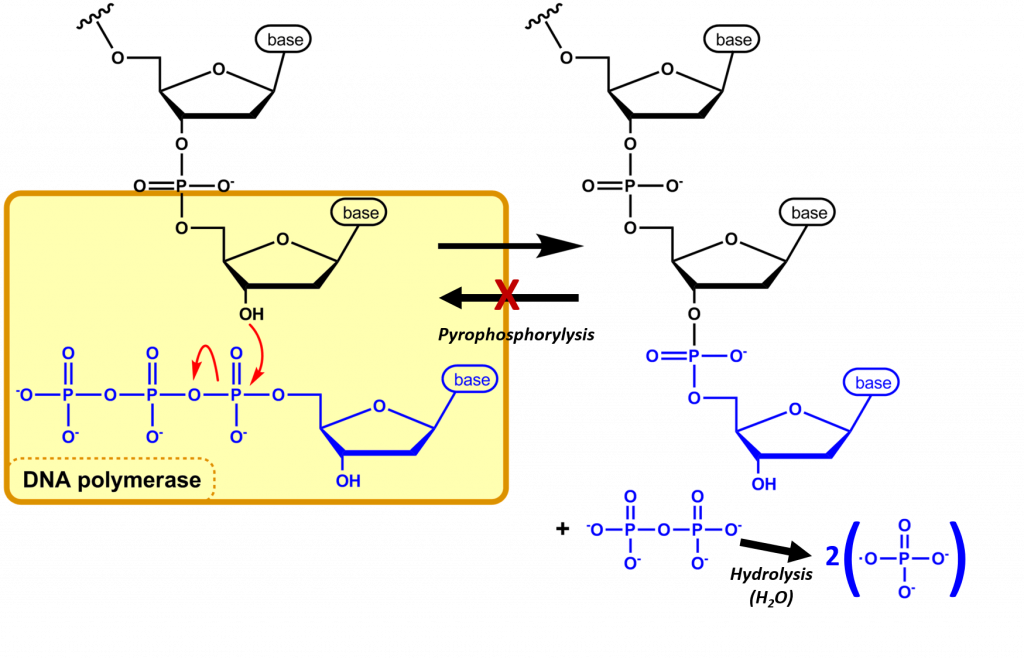
The nucleotide that is required as the monomer for the synthesis of both DNA and RNA is the high energy nucleoside triphosphate. During the incorporation of the nucleotide into the polymeric structure, two phosphate groups, (Pi-Pi , called pyrophosphate) from each triphosphate are cleaved from the incoming nucleotide and further hydrolyzed during the reaction, leaving a nucleoside monophosphate that is incorporated into the growing RNA or DNA chain (Figure 4.4). Incorporation of the incoming nucleoside triphosphate is mediated by the nucleophilic attack of the 3′-OH of the growing DNA polymer. Thus, DNA synthesis is directional, only occurring at the 3′-end of the molecule.
The further hydolysis of the pyrophosphate (Pi-Pi) releases a large amount of energy ensuring that the overall reaction has a negative ΔG. Hydrolysis of Pi-Pi –> 2Pi has a ΔG = -7 kcal/mol and is essential to provide the overall negative ΔG (-6.5 kcal/mol) of the DNA synthesis reaction. Hydrolysis of the pyrophosphate also ensures that the reverse reaction, pyrophsophorylysis, will not take place removing the newly incorporated nucleotide from the growing DNA chain.
This reaction is mediated in DNA by a family of enzymes known as DNA polymerases. Similarly, RNA polymerases are required for RNA synthesis. A more detailed description of polymerase reaction mechanisms will be covered in Chapters X and Y, covering DNA Replication and Repair, and DNA Transcription.
Figure 4.4 Nucleic Acid Synthesis: In nucleic acid synthesis, the 3’ OH of a growing chain of nucleotides attacks the α-phosphate on the next NTP to be incorporated (blue), resulting in a phosphodiester linkage and the release of pyrophosphate (PPi). The DNA polymerase further mediates the hydrolysis of the pyrophosphate preventing the reverse reaction from occurring and releasing enough energy to drive the reaction forward. The synthesis of DNA is shown in this diagram.
Image modified from Michal Sobkowski
DNA was first isolated by Friedrich Miescher in 1869. The double-helix model of DNA structure was first published in the journal Nature by James Watson and Francis Crick in 1953,(X,Y,Z coordinates in 1954) based upon the crucial X-ray diffraction image of DNA from Rosalind Franklin in 1952, followed by her more clarified DNA image with Raymond Gosling, Maurice Wilkins, Alexander Stokes, and Herbert Wilson, and base-pairing chemical and biochemical information by Erwin Chargaff. The prior model was triple-stranded DNA.
The realization that the structure of DNA is that of a double-helix elucidated the mechanism of base pairing by which genetic information is stored and copied in living organisms and is widely considered one of the most important scientific discoveries of the 20th century. Crick, Wilkins, and Watson each received one third of the 1962 Nobel Prize in Physiology or Medicine for their contributions to the discovery. (Franklin, whose breakthrough X-ray diffraction data was used to formulate the DNA structure, died in 1958, and thus was ineligible to be nominated for a Nobel Prize.)
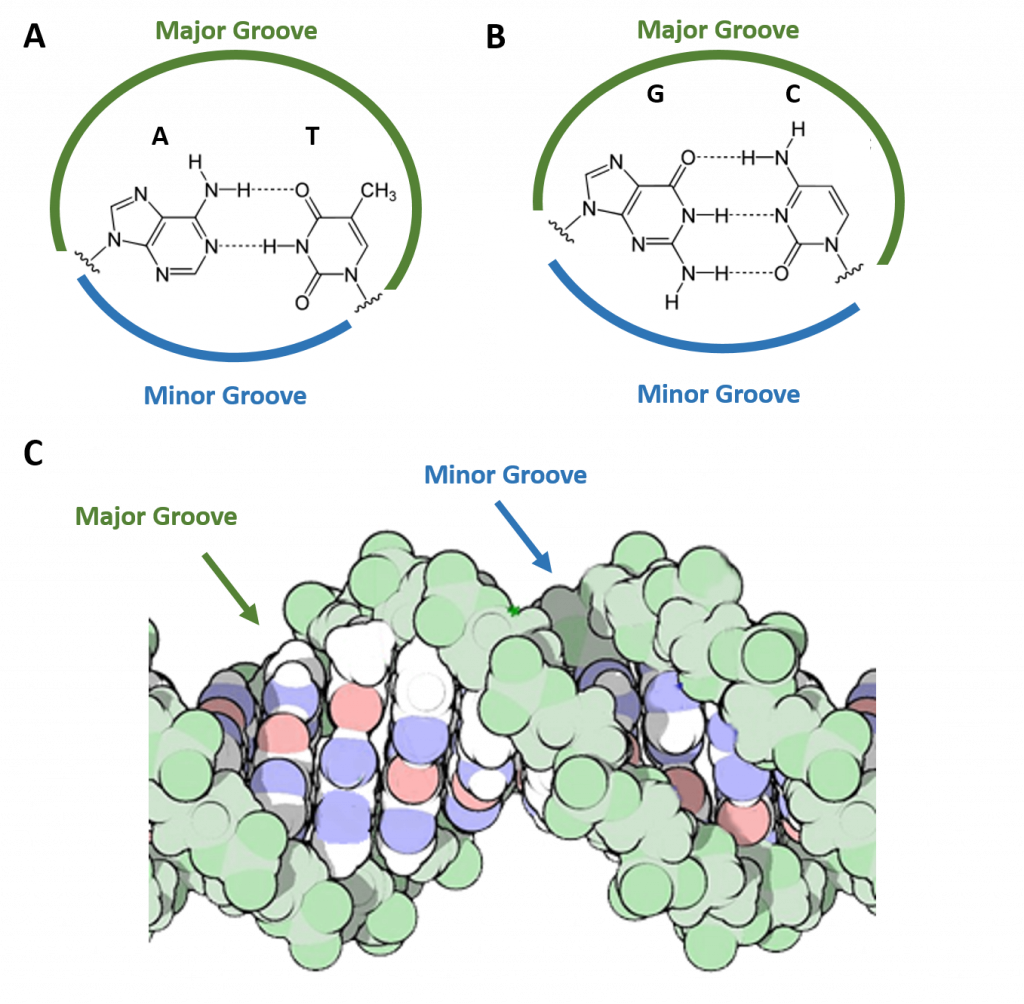
Watson and Crick proposed two strands of DNA – each in a right-hand helix – wound around the same axis. The two strands held together by H-bonding between the complimentary base pairs (A pairs with T and G pairs with C) (Figure 4.5). Note that when looking from the top view, down on a DNA base pair, that the position where the base pairs attach to the DNA backbone is not equidistant, but that attachment favors one side over the other. This creates unequal gaps or spaces in the DNA known as the major groove for the larger gap, and the minor groove for the smaller gap (Figure 4.5). Based on the DNA sequence within the region, the hydrogen-bond potential created by the nitrogen and oxygen atoms present in the nitrogenous base pairs cause unique recognition features within the major and minor grooves, allowing for specific protein recognition sites to be created.
Figure 4.5 The Major and Minor Grooves of DNA. Top view of an (A) A-T base pair and a (B) G-C base pair showing the formation of the major and minor groove sides of the DNA. (C) Side view of the DNA double helix with the major and minor grooves indicated. The DNA backbone is shown in green, potential nitrogen hydrogen-bonding locations are indicated in blue, and oxygen hydrogen-bonding locations in red.
Figure C modified from: dullhunk
back to the top
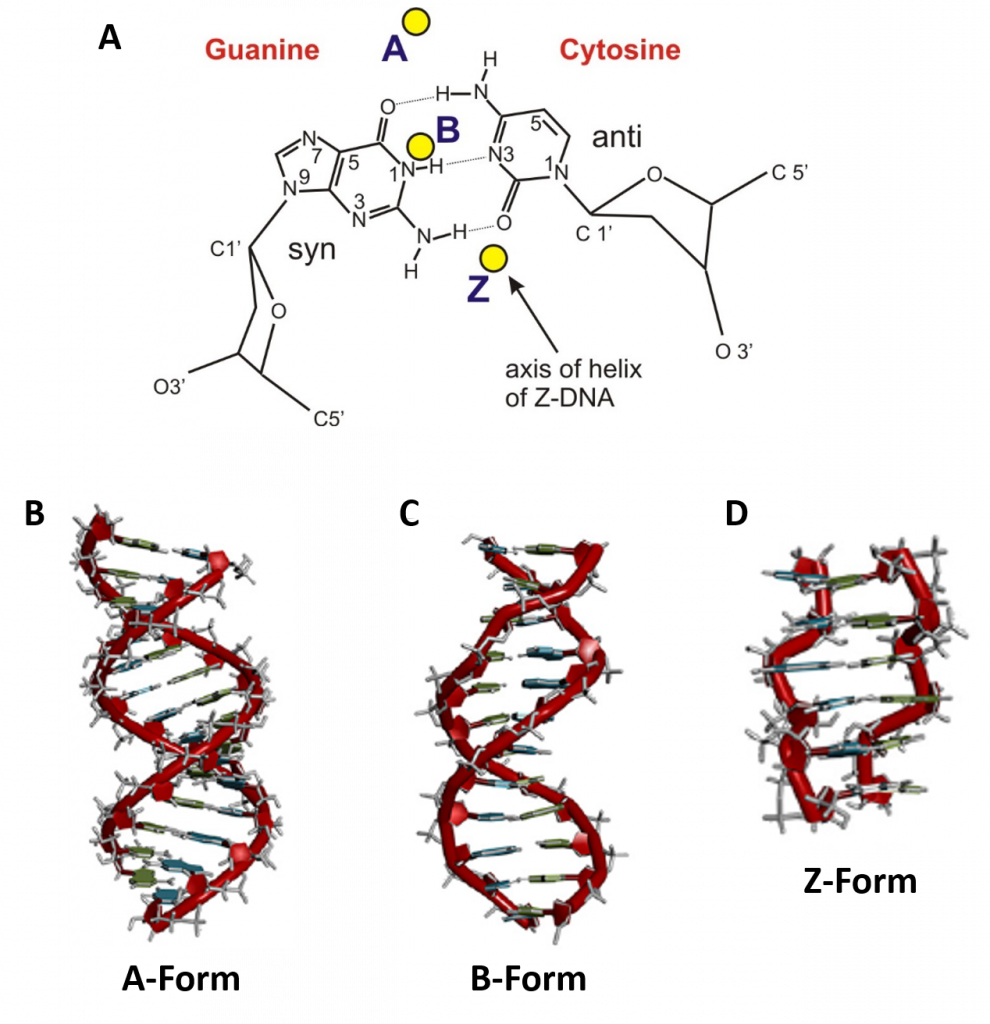
In addition to the major and minor grooves providing variation within the double helix structure, the axis alignment of the helix along with other influencing factors such as the degree of solvation, can give rise to three forms of the double helix, the A-form (A-DNA), the B-form (B-DNA), and the Z-form (Z-DNA) (Figure 4.6). Both the A- and B-forms of the double helix are right-handed spirals, with the B-form being the predominant form found in vivo. The A-form helix arises when conditions of dehydration below 75% of normal occur and have mainly been observed in vitro during X-ray crystallography experiments when the DNA helix has become dessicated. However, the A-form of the double helix can occur in vivo when RNA adopts a double stranded conformation, or when RNA-DNA complexes form. The 2′-OH group of the ribose sugar backbone in the RNA molecule prevents the RNA-DNA hybrid from adopting the B-conformation due to steric hindrance.
The third type of double helix formed is a left-handed helical structure known as the Z-form, or Z-DNA. Within this structural motif, the phosphates within the backbone appear to zigzag, providing the name Z-DNA (Figure 4.6). In vitro, the Z-form of DNA is adopted in short sequences that alternate pyrimidine and purines and when high salinity is present. However, the Z-form has been identified in vivo, within short regions of the DNA, showing that DNA is quite flexible and can adopt a variety of conformations. A comparison of features between A-, B- and Z-form DNA is shown in Table 4.1.
Dimensions of B-form (the most common) of DNA
- 0.34 nm between bp, 3.4 nm per turn, about 10 bp per turn
- 1.9 nm (about 2.0 nm or 20 Angstroms) in diameter
Figure 4.6 Major Conformations of the DNA Double Helix. (A) Shows from the top view, the different locations of the central axis in the different major forms of DNA, with the base pairs represented in the B-conformation. Side view of (B) A-form DNA, (C) B-form DNA, and (D) Z-form DNA.
Image A from: Lankenau
Images B-D from: Börner, et al (2016) Coordination Chemistry Reviews 327 DOI 10.1016/j.ccr.2016.06.002
| B-Form | A-Form | Z-Form | |
|---|---|---|---|
| helix sense | Right Handed | Right Handed | Left Handed |
| base pairs per turn | 10 | 11 | 12 |
| vertical rise per bp | 3.4 Å | 2.56 Å | 19 Å |
| rotation per bp | +36° | +33° | -30° |
| helical diameter | 19 Å | 19 Å | 19 Å |
The double stranded helix of DNA is not always stable. This is because the stair step links between the strands are noncovalent, reversible interactions. Depending on the DNA sequence, denaturation (melting) can be local or widespread and enables various crucial cellular processes to take place, including DNA replication, transcription, and repair.
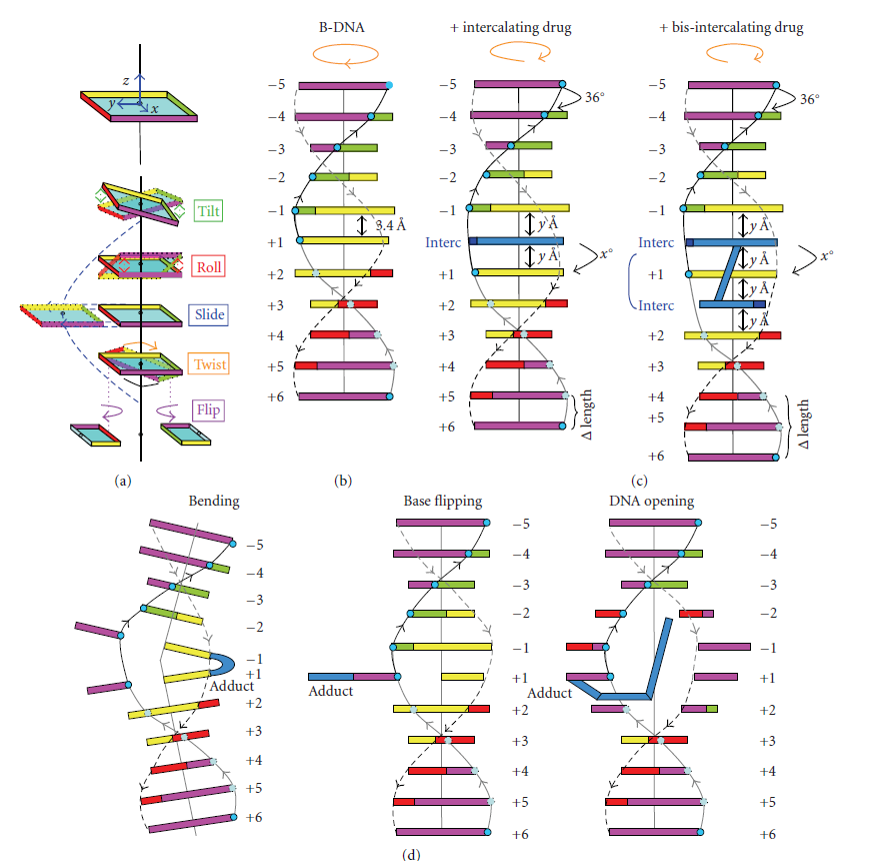
Both sequence specificity and interaction (whether covalent or not) with a small compound or a protein can induce tilt, roll and twist effects that rotate the base pairs in the x, y, or z axis, respectively (Figure 4.7), and can therefore change the helix’s overall organization. Furthermore, slide or flip effects can also modify the geometrical orientation of the helix (Figure 4.7). Hence the flip effects, and (to a lesser extent) the other above-defined movements modulate the double-strand stability within the helix or at its ends. Indeed, under physiological conditions, local DNA ‘breathing’ has been evidenced at both ends of the DNA helix and B- to Z-DNA structural transitions have been observed in internal DNA regions. These types of locally open DNA structures are good substrates for specific proteins which can also induce the opening of a ‘closed’ helix. The processes of DNA replication and repair will be discussed in more detail in Chapter XX and DNA transcription and transcriptional regulation in Chapters X and Y.
Figure 4.7 Localized Structural Modification of the DNA Double Helix. (a) Base pair orientation with x, y, and z azes result in different kinds of roatation (tilt, roll or twist) or slipping of the bases (slide, flip) regarding to the helix central axis. (b) Matove B-DNA with nearly 11 base pairs within one helical turn. (c) Mono- or bis-intercalation of a small molecule (shown in blue) between adjacent base pairs resulting in an unwinding of the DNA helix (orange arrow on the top) and a lengthening of the DNA helix (ΔLength) depending on the X and y Å values that are specific for a defined DNA intercalating compound. (d) Representation of the DNA bending, base flipping, or double strand opeing induced by some DNA destabilizing alkylating agents (adducts shown in blue). Adapted from Calladine and Drew’s schematic box representation.
Image from: Lenglet and David-Cordonnier (2010) Journal of Nucleic Acids, http://dx.doi.org/10.4061/2010/290935
back to the top
4.2 Chromosomes and Packaging
Within eukaryotic cells, DNA is organized into long linear structures called chromosomes (Figure 4.8). A chromosome is a deoxyribonucleic acid (DNA) molecule with part or all of the genetic material (genome) of an organism. Most eukaryotic chromosomes include packaging proteins which, aided by chaperone proteins, bind to and condense the DNA molecule to prevent it from becoming an unmanageable tangle. Before typical cell division, these chromosomes are duplicated in the process of DNA replication, providing a complete set of chromosomes for each daughter cell. The replicated arms of a chromosome are called chromatids. Before being separated into the daughter cells during mitosis, replicated chromatids are held together by a chromosomal structure called the centromere.

Figure 4.8 Diagram of Replicated and Condensed Eukaryotic Chromosome. (1) Chromatid – one of the two identical parts of the chromosome after S phase. (2) Centromere – the point where the two chromatids are joined together. (3) Short arm is termed p; Long arm is termed q
Image by: Magnus Manske, Dietzel65, and Tryphon
Eukaryotic organisms (animals, plants, fungi and protists) store most of their DNA inside the cell nucleus as linear nuclear DNA, and some in the mitochondria as circular mitochondrial DNA or in chloroplasts as circular chloroplast DNA. In contrast, prokaryotes (bacteria and archaea) do not have organelle structures and thus, store their DNA only in a region of the cytoplasm known as the nucleoid region. Prokaryotic chromosomes consist of double–stranded circular DNA.
The genome of a cell is often significantly larger than the cell itself. For example, if the DNA from a human cell containing 46 chromosomes were stretched out in a line, it would extend more that 6 feet (2 meters)! How is it possible that the genetic information not only fits into the cell, but fits into the cell nucleus? Eukaryota solves this problem by a combination of supercoiling and packaging DNA around the histone family of proteins (described below). Prokaryotes do not contain histones (with a few exceptions). Prokaryotes tend to compress their DNA using nucleoid-associated-proteins (NAPs) and supercoiling (Figure 4.9).
DNA supercoiling refers to the over- or under-winding of a DNA strand, and is an expression of the strain on that strand (Figure 4.9). Supercoiling is important in a number of biological processes, such as compacting DNA, and by regulating access to the genetic code. DNA supercoiling strongly affects DNA metabolism and possibly gene expression. Additionally, certain enzymes such as topoisomerases are able to change DNA topology to facilitate functions such as DNA replication or transcription.
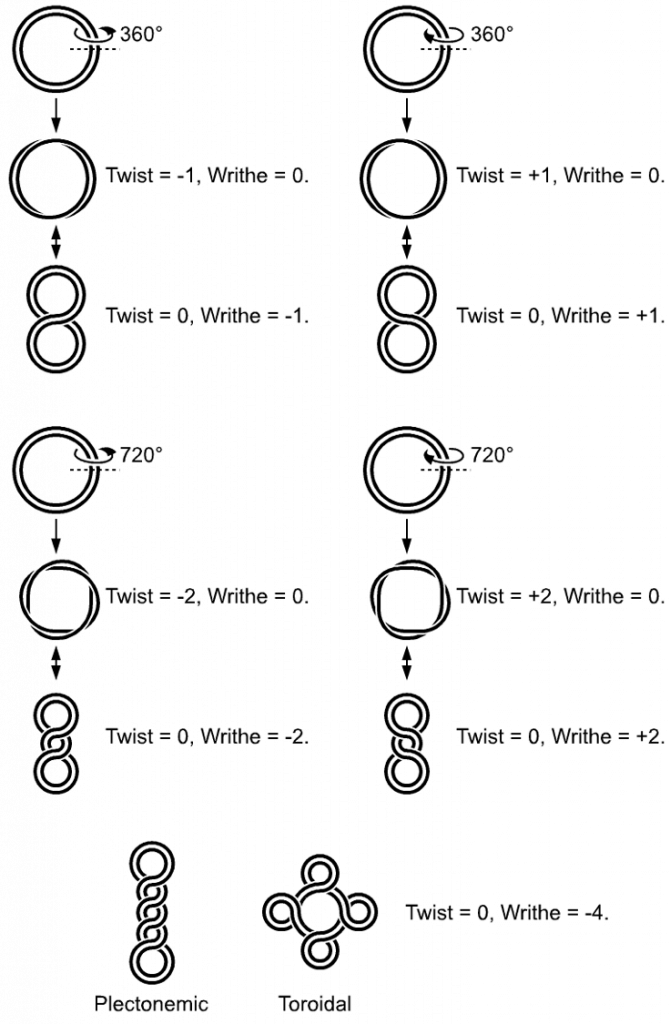
In a “relaxed” double-helical segment of B-DNA, the two strands twist around the helical axis once every 10.4–10.5 base pairs of sequence. Adding or subtracting twists, as some enzymes can do, imposes strain. If a DNA segment under twist strain were closed into a circle by joining its two ends and then allowed to move freely, the circular DNA would contort into a new shape, such as a simple figure-eight (Figure 4.9). Such a contortion is a supercoil. The noun form “supercoil” is often used in the context of DNA topology.
Figure 4.9 DNA Supercoiling. The supercoiled structure of linear DNA molecules with constrained ends. The helical nature of the DNA duplex is omitted for clarity.
Image by: Richard Wheeler
Positively supercoiled (overwound) DNA is transiently generated during DNA replication and transcription, and, if not promptly relaxed, inhibits (regulates) these processes. The simple figure eight is the simplest supercoil, and is the shape a circular DNA assumes to accommodate one too many or one too few helical twists. The two lobes of the figure eight will appear rotated either clockwise or counterclockwise with respect to one another, depending on whether the helix is over- or underwound. For each additional helical twist being accommodated, the lobes will show one more rotation about their axis. As a general rule, the DNA of most organisms is negatively supercoiled.
Lobal contortions of a circular DNA, such as the rotation of the figure-eight lobes above, are referred to as writhe. The above example illustrates that twist and writhe are interconvertible. Supercoiling can be represented mathematically by the sum of twist and writhe (Figure 4.9). The twist is the number of helical turns in the DNA and the writhe is the number of times the double helix crosses over on itself (these are the supercoils). Extra helical twists are positive and lead to positive supercoiling, while subtractive twisting causes negative supercoiling. Many topoisomerase enzymes sense supercoiling and either generate or dissipate it as they change DNA topology.
In part because chromosomes may be very large, segments in the middle may act as if their ends are anchored. As a result, they may be unable to distribute excess twist to the rest of the chromosome or to absorb twist to recover from underwinding—the segments may become supercoiled, in other words. In response to supercoiling, they will assume an amount of writhe, just as if their ends were joined.
Supercoiled circular DNA forms two major structures; a plectoneme or a toroid, or a combination of both (Figure 4.9). A negatively supercoiled DNA molecule will produce either a one-start left-handed helix, the toroid, or a two-start right-handed helix with terminal loops, the plectoneme. Plectonemes are typically more common in nature, and this is the shape most bacterial plasmids will take (Figure 4.10). For larger molecules it is common for hybrid structures to form – a loop on a toroid can extend into a plectoneme (Figure 4.10). DNA supercoiling is an important for DNA packaging within all cells, and seems to also play a role in gene expression.
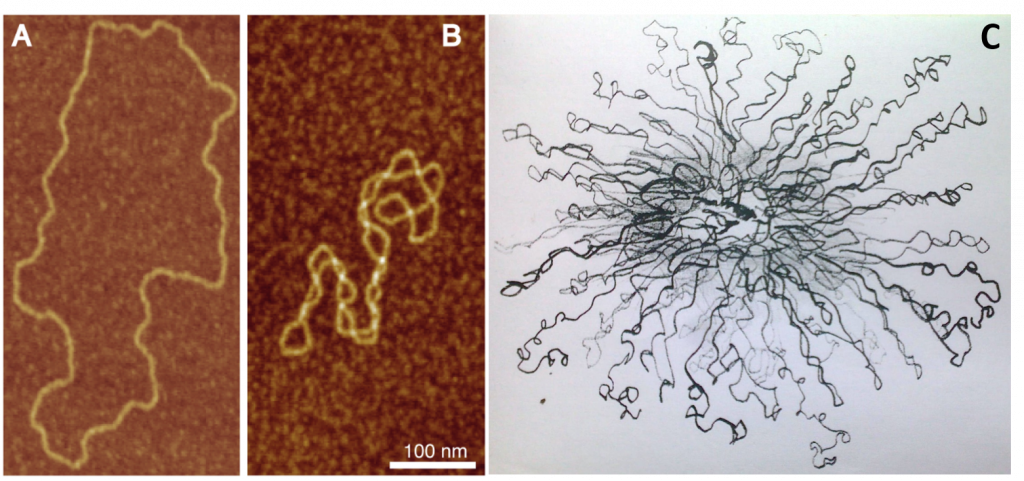
Figure 4.10 Bacterial DNA Supercoiling. Atomic force microscopy (AFM) visualization of torsionally relaxed (A), and negativey supercoiled (B) bacterial plasmids pBR322. (C) Electron microscopy image of the E. coli chromosomal DNA displaying a hybrid toroidal-plectoneme structure.
Image A and B from: Witz, G. and Stasiak, A. (2009) Nucleic Acids Research 38(7):2119-2133.
Image C from: Prokaryotic Chromosomes
In addition to forming supercoiled structure, circular chromosomes from bacteria have been shown to undergo the processes of catenation and knotting upon the inhibition of topoisomerase enzymes. Catenation is the process by which two circular DNA strands are linked together like chain links, whereas DNA knotting is the interlooping structures occurring within a single circular DNA structure. In vivo, the action of topoisomerase enzymes is critical to keep knots and catenoids from tangling the DNA structure.

Figure 4.11 DNA Catenation and Knotting. Upper structure shows the negative supercoiled form of bacterial DNA. The inhibition of topoisomerase enzyme activity leads to the relaxation, catenation and knotting of the chromosomal structure.
Image from: Harms, A. et al. (2015) Cell Reports 12(9):1497-1507.
Note the circular nature of chloroplast and mitochondrial DNA, suggesting a bacterial origin for both of these organelle structures. Sequence alignments further lend support for the endosymbiotic theory, which proposes that bacteria were engulfed by early eukaryotic organisms and subsequently became symbiotic to their eukaryotic counterpart, rather than being digested.
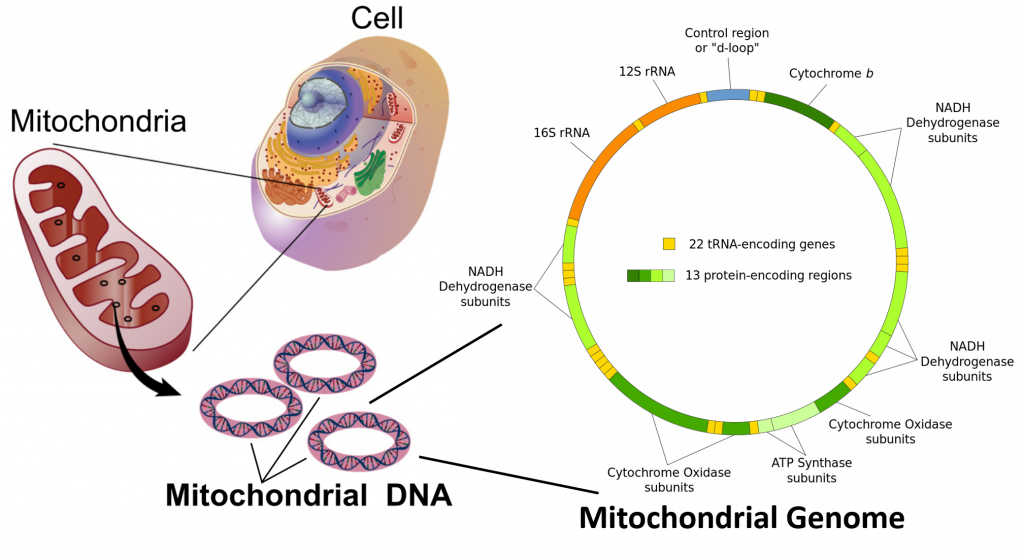
In the cells of extant organisms, the vast majority of the proteins present in the mitochondria (numbering approximately 1500 different types in mammals) are coded for by nuclear DNA. However, sequencing of the human mitochondrial genome has revealed 16,569 base pairs encoding for 13 proteins (Figure 4.12). Many of the mitochondrially produced proteins are required for electron transport during the production of ATP (Figure 4.12).
Figure 4.12 Mitochondrial Genome. Mitochondria are organelle structures containing a double membrane, thought to have originated as an independent prokaryotic organism that was originally engulfed by a eukaryotic organism, where it became a symbiotic counterpart. Mitochondria contain circular chromosomal DNA that shares high sequence similarity with alphaprotobacteria. The human mitochondrial genome contains 16,569 base pairs encoding for 13 proteins and ribosomal RNA (rRNA) components.
Images adapted from: The National Human Genome Research Institute and Shanel, Knopfkind, and JHC.
back to the top
Within eukaryotic chromosomes, chromatin proteins, known as histones, compact and organize DNA. These compacting structures guide the interactions between DNA and other proteins, helping control which parts of the DNA are transcribed.
Histones are highly alkaline proteins found in eukaryotic cell nuclei that package and order the DNA into structural units called nucleosomes. They are the chief protein components of chromatin, acting as spools around which DNA winds, and playing a role in gene regulation. Without histones, the unwound DNA in chromosomes would be very long (a length to width ratio of more than 10 million to 1 in human DNA). For example, each human diploid cell (containing 23 pairs of chromosomes) has about 1.8 meters of DNA; wound on the histones, the diploid cell has about 90 micrometers (0.09 mm) of chromatin.
Five major families of histones exist: H1/H5, H2A, H2B, H3, and H4. Histones H2A, H2B, H3 and H4 are known as the core histones, while histones H1/H5 are known as the linker histones.
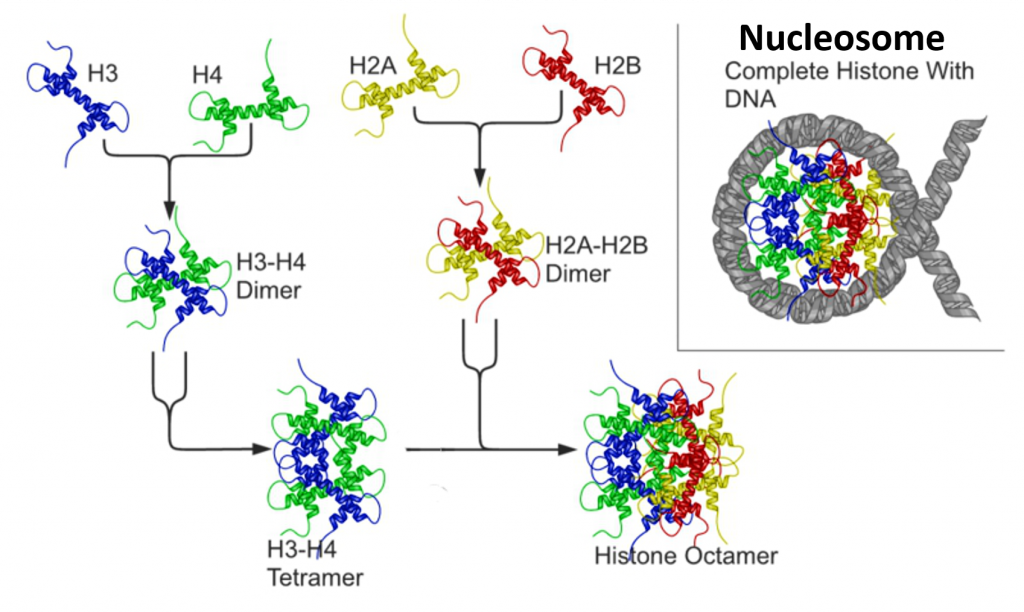
The core histones all exist as dimers, which are similar in that they all possess the histone fold domain: three alpha helices linked by two loops (Figure 4.13). It is this helical structure that allows for interaction between distinct dimers, particularly in a head-tail fashion (also called the handshake motif). The resulting four distinct dimers then come together to form one octameric nucleosome core, approximately 63 Angstroms in diameter. Around 146 base pairs (bp) of DNA wrap around this core particle 1.65 times in a left-handed super-helical turn to give a particle of around 100 Angstroms across, called a nucleosome.
Figure 4.13 Nucleosome Core Structure. Histones H2A and H2B dimerize, and Histones H3 and H4 dimerize. Two dimers of each join to form a histone core octomer. The DNA double helix winds 1.65 times around the octomer core forming the nucleosome structure.
Image adapted from: Nucleosome Structure
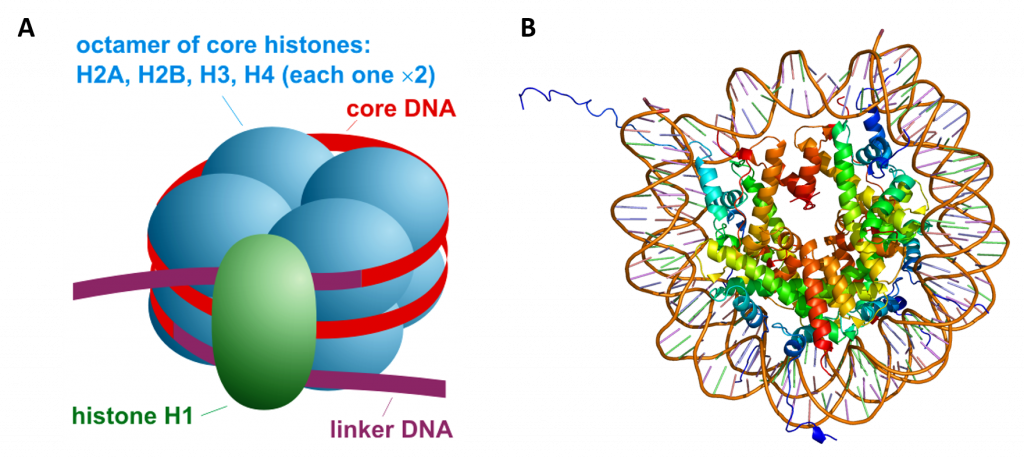
The linker histone H1 binds the nucleosome at the entry and exit sites of the DNA, thus locking the DNA into place and allowing the formation of higher order structure (Figure 4.14). The most basic such formation is the 10 nm fiber or beads on a string conformation. This involves the wrapping of DNA around nucleosomes with approximately 50 base pairs of DNA separating each pair of nucleosomes (also referred to as linker DNA).
The nucleosome contains over 120 direct protein-DNA interactions and several hundred water-mediated ones. Direct protein – DNA interactions are not spread evenly about the octamer surface but rather located at discrete sites. These are due to the formation of two types of DNA binding sites within the octamer; the α1α1 site, which uses the α1 helix from two adjacent histones, and the L1L2 site formed by the L1 and L2 loops. Salt links and hydrogen bonding between both side-chain basic and hydroxyl groups and main-chain amides with the DNA backbone phosphates form the bulk of interactions with the DNA. This is important, given that the ubiquitous distribution of nucleosomes along genomes requires it to be a non-sequence-specific DNA-binding factor. Although nucleosomes tend to prefer some DNA sequences over others, they are capable of binding practically to any sequence, which is thought to be due to the flexibility in the formation of these water-mediated interactions. In addition, non-polar interactions are made between protein side-chains and the deoxyribose groups, and an arginine side-chain intercalates into the DNA minor groove at all 14 sites where it faces the octamer surface. The distribution and strength of DNA-binding sites about the octamer surface distorts the DNA within the nucleosome core. The DNA is non-uniformly bent and also contains twist defects. The twist of free B-form DNA in solution is 10.5 bp per turn. However, the overall twist of nucleosomal DNA is only 10.2 bp per turn, varying from a value of 9.4 to 10.9 bp per turn.
The histone tail extensions constitute up to 30% by mass of histones, but are not visible in the crystal structures of nucleosomes due to their high intrinsic flexibility, and have been thought to be largely unstructured (Figure 4.14). The N-terminal tails of histones H3 and H2B pass through a channel formed by the minor grooves of the two DNA strands, protruding from the DNA every 20 bp. The N-terminal tail of histone H4, on the other hand, has a region of highly basic amino acids (16-25), which, in the crystal structure, forms an interaction with the highly acidic surface region of a H2A-H2B dimer of another nucleosome, being potentially relevant for the higher-order structure of nucleosomes. This interaction is thought to occur under physiological conditions also, and suggests that acetylation of the H4 tail distorts the higher-order structure of chromatin.
Figure 4.14 Overall Nucleosome Structure. (A) Side view diagram of the nucleosome structure with the histone octomer shown in blue, the DNA double helix in red, and the histone H1 linker in green. (B) Shows a top view rendering of the histone octomer with the associated DNA helix. Note that the Histone tails from H3 and H2B protude from the DNA.
Image A from: Darekk2 Image B from: EMW
The formation of the DNA double helix represents the first order packaging of the chromosome structure (Figure 4.15). The formation of nucleosomes represent the second level of packaging for eukaryotic chromosomes. In vitro data suggests that nucleosomes are then arranged into either a solenoid structure which consists of 6 nucleosomes linked together by the Histone H1 linker proteins or a zigzag structure that is similar to the solenoid construct (Figure 4.15). Both the solenoid and zigzag structures are approximately 30 nm in diamater. The solenoid and zigzag structures reported from in vitro data have not yet been confirmed to occur in vivo.
During interphase, each chromosome occupies a spatially limited, roughly elliptical domain which is known as a chromosome territory (CT). Each chromosome territory is comprised of higher order chromatin units of ~1 Mb each. These units are likely built up from smaller loop domains that contain the solenoid/zigzag structural motifs. On the other hand, 1Mb domains can themselves serve as smaller units in higher-order chromatin structures.
Chromosome territories are known to be arranged radially around the nucleus. This arrangement is both cell and tissue-type specific and is also evolutionary conserved. The radial organization of chromosome territories was shown to correlate with their gene density and size. In this case, the gene-rich chromosomes occupy interior positions, whereas larger, gene-poor chromosomes, tend to be located around the periphery. Chromosome territories are also dynamic structures, with genes able to relocate from the periphery towards the interior once they have been ‘switched on’. In other cases, genes may move in the opposite direction, or simply maintain their position. The eviction of genes from their chromosome territories into the interchromatin compartment or a neighboring chromosome territory is often accompanied by the formation of large decondensed chromatin loops.

Figure 4.15 Chromosome Structure. (1) DNA double helix is approximately 2 nm in diameter. (2) The nucleosome core structure is approximately 11 nm in diameter. (3) The solenoid/zigzag structure is approximately 30 nm in diameter and is proposed to form chromosome loops (4) during cellular interphase and more condensed chromosome territories (5) during mitosis.
Image by: MBInfo
Models describing chromosome territory arrangement
With the development of high-throughput biochemical techniques, such as 3C (‘chromosome conformation capture’) and 4C (‘chromosome conformation capture-on-chip’ and ‘circular chromosome conformation capture’), numerous spatial interactions between neighbouring chromatin territories have been described (Figure 4.16). These descriptions have been supplemented with the construction of spatial proximity maps for the entire genome (e.g., for a human lymphoblastoid cell line). Together, these observations and physical simulations have led to the proposal of various models that aim to define the structural organization of chromosome territories:

Figure 4.16 Computer Models of Chromosome Territory (CT) Structure. On the CT-IC model, the space between discrete CTs can be visualized in light and electron microscope and is called interchromatin compartment (IC). Transcription factories (TF, green color) are localized predominantly in perichromatin region. In the ICN model, interchromatin compartment is not apparent. Instead, the space between CTs is occupied by intermingling decondensed chromatin loops, which often share the same transcription factories.
Image by: MBInfo
1. The chromosome territory-interchromatin compartment (CT-IC) model describes two principal compartments: chromosome territories (CTs) and an interchromatin compartment (IC). In this model, chromosome territories build up an interconnected chromatin network that is associated with an adjacent 3D space called the interchromatin compartment. The latter can be observed using both light and electron microscopy.
Within a single chromosome territory, the interphase chromosome is divided into defined regions based on the level of chromosome condensation. Here, the inner part of the interphase chromosome is comprised of more condensed chromatin domains or higher-order chromatin fibers, while a thin (<200 nm) layer of more decondensed chromatin, known as the perichromatin region, can be found around the chromosomal periphery. Functionally, the perichromatin region represents the major transcriptional compartment, and is also the region where most co-transcriptional RNA splicing takes place. DNA replication [20] and DNA repair [21] is also predominately carried out within the perichromatin region. Finally, nascent RNA transcripts, referred to as perichromatin fibrils, are also generated in the perichromatin region. Perichromatin fibrils are then subjected to the splicing events by the factors, provided from the interchromatin compartment.
The lattice model, proposed by Dehgani et al. is based on reports that transcription also occurs within the inner, more condensed chromosome territories and not only at the interface between the interchromatin compartment and the perichromatin region. Using ESI (electron spectroscopic imaging), Dehgani et al. showed that chromatin was organized as an array of deoxy-ribonucleoprotein fibers of 10–30 nm in diameter. In this study, the interchromatin compartments, which are described in the CT-IC model as large channels between chromosome territories, were not apparent. Instead, chromatin fibers created a loose meshwork of chromatin throughout the nucleus that intermingled at the periphery of chromosome territories. Thus, inter- and intra-chromosomal spaces within this meshwork are essentially contiguous and together form the intra-nuclear space.
2. The interchromatin network (ICN) model predicts that intermingling chromatin fibers/loops can make both cis- (within the same chromosome) and trans- (between different chromosomes) contacts. This intermingling is uniform and makes distinction between the chromosome territory and interchromatin compartment functionally meaningless. The advantage of the ICN model is that it permits high chromatin dynamics and diffusion-like movements. The authors propose that ongoing transcription influences the degree of intermingling between specific chromosomes by stabilizing associations between particular loci. Such interactions are likely to depend on the transcriptional activity of the loci, and are therefore cell-type specific.
The cell type-specific organization of chromosome territories has been studied by measuring the volume and frequency of intermingling between heterologous chromosomes. By using 3C (chromosome conformation capture) and FISH (fluorescence in situ hybridization) to map the regions of chromosome intermingling, it was revealed that these regions contain a higher density of active genes and are enriched with markers of transcriptional activation and repression, such as activated RNAPII. By comparing the positions of the CTs in undifferentiated mouse embryonic stem (ES) cells, ES cells in early stages of differentiation, and terminally differentiated NIH3T3 cells, it was shown that fully differentiated cells had a higher enrichment of RNAPII, compared to undifferentiated or less-differentiated cells. The findings support the notion that the intermingling regions have functional significance in the nucleus and provide a basis for understanding how the radial and relative positions of chromosomal territories evolve during the process of differentiation, explaining their organization in a cell type-dependent manner.
3. The Fraser and Bickmore model emphasizes the functional importance of giant chromatin loops, which originate from chromosome territories and expand across the nuclear space in order to share transcription factories. In this case, both cis- and trans- loops of decondensed chromatin can be co-expressed and co-regulated by the same transcription factory.
4. The Chromatin polymer models assume a broad range of chromatin loop sizes and predict the observed distances between genomic loci and chromosome territories, as well as the probabilities of contacts being formed between given loci. These models apply physics-based approaches that highlight the importance of entropy for understanding nuclear organization. By proposing the existence of conformational chromatin ensembles with structures based on three possible homopolymer states, these models also provide alternative structures to the traditional 30 nm chromatin fiber, which has been brought into question following recent studies.
With a lack of experimental evidence to support these described models, it must be remembered that they serve only to hypothesize the structural and chemical properties of intermediate chromatin structures, and to highlight unanswered questions. For example, the mechanisms that exist to control the rate and the extent of chromatin movement remain to be defined
back to the top
Telomere Structures
At the ends of the linear chromosomes are specialized regions of DNA called telomeres (Figure 4.17). The main function of these regions is to allow the cell to replicate chromosome ends using the enzyme telomerase, as the enzymes that normally replicate DNA cannot copy the extreme 3′ ends of chromosomes. These specialized chromosome caps also help protect the DNA ends, and stop the DNA repair systems in the cell from treating them as damage to be corrected. In human cells, telomeres are usually lengths of single-stranded DNA containing several thousand repeats of a simple TTAGGG sequence.
During DNA replication, the double stranded DNA is unwound and DNA polymerase synthesizes new strands. However, as DNA polymerase moves in a unidirectional manner (from 5’ to 3’), only the leading strand can be replicated continuously. In the case of the lagging strand, DNA replication is discontinuous. In humans small RNA primers attach to the lagging strand DNA, and the DNA is synthesized in small stretches of about 100-200 nucleotides, which are termed Okazaki fragments. The RNA primers are removed, replaced with DNA and the Okazaki fragments ligated together. At the end of the lagging strand, it is impossible to attach an RNA primer, meaning that there will be a small amount of DNA lost each time the cell divides. This ‘end replication problem’ has serious consequences for the cell as it means the DNA sequence cannot be replicated correctly, with the loss of genetic information.
In order to prevent this, telomeres are repeated hundreds to thousands of times at the end of the chromosomes. Each time cell division occurs, a small section of telomeric sequences are lost to the end replication problem, thereby protecting the genetic information. At some point, the telomeres become critically short. This attrition leads to cell senescence, where the cell is unable to divide, or apoptotic cell death. Telomeres are the basis for the Hayflick limit, the number of times a cell is able to divide before reaching senescence.
Telomeres can be restored by the enzyme telomerase, which extends telomeres length (Figure 4.17). Telomerase activity is found in cells that undergo regular division, such as stem cells and lymphocyte cells of the immune system. Telomeres can also be extended through the Alternative Lengthening of Telomeres (ALT) pathway. In this case, rather than being extended, telomeres are switched between chromosomes by homologous recombination. As a result of the telomere swap, one set of daughter cells will have shorter telomeres, and the other set will have longer telomeres.
A downside to telomere extension is the potential for uncontrolled cell division and cancer. Abnormally high telomerase activity has been found in the majority of cancer cells, and non-telomerase tumors often exhibit ALT pathway activation. As well as the potential for losing genetic information, cells with short telomeres are at a high risk for improper chromosome recombination, which can lead to genetic instability and aneuploidy (an abnormal number of chromosomes).
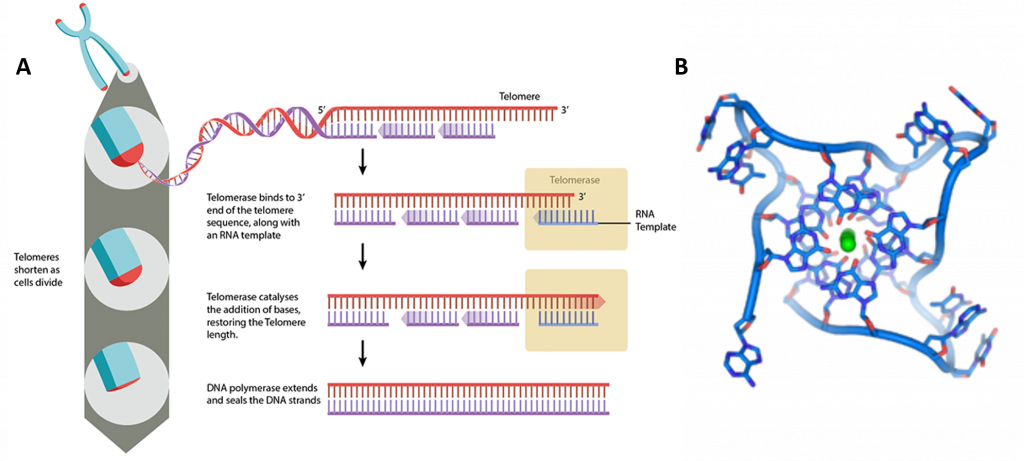
Figure 4.17 Telomere Structure. (A) Telomeres are located at the end of chromosomes, where they help protect against the loss of DNA during replication. (B) DNA quadruplex formed by telomere repeats. The looped conformation of the DNA backbone is very different from the typical DNA helix. The green spheres in the center represent potassium ions.
Image (A) by: MBInfo and Image (B) by: Thomas Splettstoesser
These guanine-rich telomere sequences may also stabilize chromosome ends by forming structures of stacked sets of four-base units, rather than the usual base pairs found in other DNA molecules (Figure 4.17). Here, four guanine bases form a flat plate and these flat four-base units then stack on top of each other, to form a stable G-quadruplex structure. These structures are stabilized by hydrogen bonding between the edges of the bases and chelation of a metal ion in the centre of each four-base unit. Other structures can also be formed, with the central set of four bases coming from either a single strand folded around the bases, or several different parallel strands, each contributing one base to the central structure.
In addition to these stacked structures, telomeres also form large loop structures called telomere loops, or T-loops. Here, the single-stranded DNA curls around in a long circle stabilized by telomere-binding proteins. At the very end of the T-loop, the single-stranded telomere DNA is held onto a region of double-stranded DNA by the telomere strand disrupting the double-helical DNA and base pairing to one of the two strands. This triple-stranded structure is called a displacement loop or D-loop.
back to the top
4.3 Sequencing the Human Genome
In humans, each cell normally contains 23 pairs of chromosomes, for a total of 46. Twenty-two of these pairs, called autosomes, and look the same in both males and females. The 23rd pair, the sex chromosomes, differ between males and females. Females have two copies of the X chromosome, while males have one X and one Y chromosome (Figure 4.18). Each species has a unique chromosomal complement. For example, chickens have 39 pairs of chromosomes (78 total) with 38 autosomal pairs and one pair of sex chromosomes (Z and W). In this species, ZW chickens are female and ZZ chickens are male.
The total length of the human genome is over 3 billion base pairs. The total length of the human genome is over 3 billion base pairs. The genome also includes the mitochondrial DNA (Figure 4.12).

Figure 4.18 DNA Karyotype. The 22 autosomes are numbered by size. The other two chromosomes, X and Y, are the sex chromosomes. This picture of the human chromosomes lined up in pairs is called a karyotype. Karyotypes are prepared using standardized staining procedures that reveal characteristic structural features for each chromosome, usually from white blood cells.
Image by: U.S. National Library of Medicine
The first human genome sequences were published in nearly complete draft form in February 2001 by the Human Genome Project and Celera Corporation. Completion of the Human Genome Project’s sequencing effort was announced in 2004 with the publication of a draft genome sequence. Researchers working on the human genome project deciphered the human genome in three major ways: determining the order, or “sequence,” of all the bases in our genome’s DNA; making maps that show the locations of genes for major sections of all our chromosomes; and producing what are called linkage maps, through which inherited traits (such as those for genetic disease) can be tracked over generations.
Prior to the acquisition of the full genome sequence, estimates of the number of human genes ranged from 50,000 to 140,000 (with occasional vagueness about whether these estimates included non-protein coding genes). As genome sequence quality and the methods for identifying protein-coding genes improved, the count of recognized protein-coding genes dropped to 19,000-20,000. However, a fuller understanding of the role played by genes expressing regulatory RNAs that do not encode proteins has raised the total number of genes to at least 46,831, plus another 2300 micro-RNA genes. By 2012, functional DNA elements that encode neither RNA nor proteins have also been noted. Protein-coding sequences account for only a very small fraction of the genome (approximately 1.5%), and the rest is associated with non-coding RNA genes, regulatory DNA sequences, long interspersed nucleotide elements (LINEs), short interspersed nucleotide elements (SINEs), introns, and sequences for which as yet no function has been determined.
Recall that a gene is defined as a sequence of nucleotides in DNA or RNA that codes for a molecule that has a function. During gene expression, the DNA is first copied into RNA. The RNA can be directly functional or be the intermediate template for a protein that performs a function. Gene structure is the organisation of specialized sequence elements within a gene (Figure 4.19). Genes contain the information necessary for living cells to survive and reproduce. The processes of transcription which leads to the production of the RNA from the DNA template, and translation which produces protein from the messenger RNA (mRNA) sequence are controlled by specific sequence elements or regions within the gene. Every gene, therefore, requires multiple sequence elements to be functional. This includes the sequence that actually encodes the functional protein or ncRNA, as well as multiple regulatory sequence regions. These regions may be as short as a few base pairs, up to many thousands of base pairs long.
Much of gene structure is broadly similar between eukaryotes and prokaryotes. These common elements largely result from the shared ancestry of cellular life in organisms with roughly 3.8 billion years of evolution. Key differences in gene structure between eukaryotes and prokaryotes reflect their divergent transcription and translation machinery. Understanding gene structure is the foundation of understanding gene annotation, expression, and function.

Figure 4.19. The Process of Eukaryotic Gene Expression. Upper blue panel shows the structural elements common to eukaryotic genes. The process of gene transcription produces a messenger RNA (mRNA) molecule that must be modified post-translationally, gray panel, to remove the non-coding intron sequences and add the 5′-CAP and Poly-A-Tail sections. The mature mRNA is transported from the nucleus to the cytoplasm where it is translated by the ribosome into the protein sequence, red panel.
Image from Wikipedia
The structures of both eukaryotic and prokaryotic genes involve several nested sequence elements. Each element has a specific function in the multi-step process of gene expression. The sequences and lengths of these elements vary, but the same general functions are present in most genes. Although DNA is a double-stranded molecule, typically only one of the strands encodes information that the RNA polymerase reads to produce protein-coding mRNA or non-coding RNA. This ‘sense’ or ‘coding’ strand, runs in the 5′ to 3′ direction where the numbers refer to the carbon atoms of the backbone’s ribose sugar. The open reading frame (ORF) of a gene is therefore usually represented as an arrow indicating the direction in which the sense strand is read.
Regulatory sequences are located at the extremities of genes. These sequence regions can either be next to the transcribed region (the promoter) or separated by many kilobases (enhancers and silencers). The promoter is located at the 5′ end of the gene and is composed of a core promoter sequence and a proximal promoter sequence. The core promoter marks the start site for transcription by binding RNA polymerase and other proteins necessary for copying DNA to RNA. The proximal promoter region binds transcription factors that modify the affinity of the core promoter for RNA polymerase. Genes may be regulated by multiple enhancer and silencer sequences that further modify the activity of promoters by binding activator or repressor proteins. Enhancers and silencers may be distantly located from the gene, many thousands of base pairs away. The binding of different transcription factors, therefore, regulates the rate of transcription initiation at different times and in different cells.
Regulatory elements can overlap one another, with a section of DNA able to interact with many competing activators and repressors as well as RNA polymerase. For example, some repressor proteins can bind to the core promoter to prevent polymerase binding. For genes with multiple regulatory sequences, the rate of transcription is the product of all of the elements combined. Binding of activators and repressors to multiple regulatory sequences has a cooperative effect on transcription initiation.
An additional layer of regulation occurs for protein coding genes after the mRNA has been processed to prepare it for translation to protein. Only the region between the start and stop codons encodes the final protein product. The flanking untranslated regions (UTRs) contain further regulatory sequences. The 3′ UTR contains a terminator sequence, which marks the endpoint for transcription and releases the RNA polymerase. The 5’ UTR binds the ribosome, which translates the protein-coding region into a string of amino acids that fold to form the final protein product. In the case of genes for non-coding RNAs the RNA is not translated but instead folds to be directly functional.
The structure of eukaryotic genes includes features not found in prokaryotes. Most of these relate to post-transcriptional modification of pre-mRNAs to produce mature mRNA ready for translation into protein. Eukaryotic genes typically have more regulatory elements to control gene expression compared to prokaryotes. This is particularly true in multicellular eukaryotes, where gene expression varies widely among different tissues.
A key feature of the structure of eukaryotic genes is that their transcripts are typically subdivided into exon and intron regions. Exon regions are the coding portion of the mRNA and are retained in the final mature mRNA molecule, while intron regions are non-coding and are spliced out (excised) during post-transcriptional processing. Indeed, the intron regions of a gene can be considerably longer than the exon regions. Once spliced together, the exons form a single continuous protein-coding region, and the splice boundaries are not detectable. Eukaryotic post-transcriptional processing also adds a 5′ cap to the start of the mRNA and a poly-adenosine tail (poly-A-tail) to the end of the mRNA. These additions stabilize the mRNA and direct its transport from the nucleus to the cytoplasm.
The overall organization of prokaryotic genes is markedly different from that of the eukaryotes. The most obvious difference is that prokaryotic ORFs are often grouped into a structure that is called a polycistronic operon under the control of a shared set of regulatory sequences (Figure 4.20). These ORFs are all transcribed onto the same mRNA and so are co-regulated and often serve related functions. Each ORF typically has its own ribosome binding site (RBS) so that ribosomes simultaneously translates the different ORFs on the same mRNA. Some operons also display translational coupling, where the translation rates of multiple ORFs within an operon are linked. This can occur when the ribosome remains attached at the end of an ORF and simply translocates along to the next without the need for a new RBS. Translational coupling is also observed when translation of an ORF affects the accessibility of the next RBS through changes in RNA secondary structure. Having multiple ORFs on a single mRNA is only possible in prokaryotes because their transcription and translation take place at the same time and in the same subcellular location.

Figure 4.20 The Process of Prokaryotic Gene Expression. Upper blue panel displays the organization of a typical prokaryotic polycistronic operon, wherein mulitiple genes are regulated by common regulatory elements and are transcribed as a single mRNA. Unlike eukaryotic systems, there is little to no post-translational modification of the resulting mRNA and protein translation, red panel, often ensues before transcription is complete.
Image from Wikipedia
The operator sequence next to the promoter is the main regulatory element in prokaryotes. Repressor proteins bound to the operator sequence physically obstructs the RNA polymerase enzyme, preventing transcription. Riboswitches are another important regulatory sequence commonly present in prokaryotic UTRs. These sequences switch between alternative secondary structures in the RNA depending on the concentration of key metabolites. The secondary structures then either block or reveal important sequence regions such as ribosomal binding sites. Introns are extremely rare in prokaryotes and therefore do not play a significant role in prokaryotic gene regulation.
Overall, the packaging, unpackaging, replication and transcription of DNA is a highly dynamic process that is constantly being moderated by signals and cues from the environment. The following video created by Drew Berry for WEHI.tv provides one of the most dynamic views of the major packaging and processing of DNA within the cell. In later chapters we will explore the processes of DNA replication and transcription in greater detail.
Video 4.1 DNA animations by wehi.tv for Science-Art exhibition. By far one of the best animations depicting DNA packaging, replication and transcription.
Created by Drew Berry and Etsuko Uno and sponsored by the Walter and Eliza Hall Institute of Medical Research
back to the top
4.4 References
Börner, R., Kowerko, D., Miserachs, H.G., Shaffer, M., and Sigel, R.K.O. (2016) Metal ion induced heterogeneity in RNA folding studied by smFRET. Coordination Chemistry Reviews 327 DOI: 10.1016/j.ccr.2016.06.002 Available at: https://www.researchgate.net/publication/303846502_Metal_ion_induced_heterogeneity_in_RNA_folding_studied_by_smFRET
Hardison, R. (2019) B-Form, A-Form, and Z-Form of DNA. Chapter in: R. Hardison’s Working with Molecular Genetics. Published by LibreTexts. Available at: https://bio.libretexts.org/Bookshelves/Genetics/Book%3A_Working_with_Molecular_Genetics_(Hardison)/Unit_I%3A_Genes%2C_Nucleic_Acids%2C_Genomes_and_Chromosomes/2%3A_Structures_of_Nucleic_Acids/2.5%3A_B-Form%2C_A-Form%2C_and_Z-Form_of_DNA
Lenglet, G., David-Cordonnier, M-H., (2010) DNA-destabilizing agents as an alternative approach for targeting DNA: Mechanisms of action and cellular consequences. Journal of Nucleic Acids 2010, Article ID: 290935, DOI: 10.4061/2010/290935 Available at: https://www.hindawi.com/journals/jna/2010/290935/
Mechanobiology Institute (2018) What are chromosomes and chromosome territories? Produced by the National University of Singapore. Available at: https://www.mechanobio.info/genome-regulation/what-are-chromosomes-and-chromosome-territories/
National Human Genome Research Institute (2019) The Human Genome Project. National Institutes of Health. Available at: https://www.genome.gov/human-genome-project
Wikipedia contributors. (2019, July 8). DNA. In Wikipedia, The Free Encyclopedia. Retrieved 02:41, July 22, 2019, from https://en.wikipedia.org/w/index.php?title=DNA&oldid=905364161
Wikipedia contributors. (2019, July 22). Chromosome. In Wikipedia, The Free Encyclopedia. Retrieved 15:18, July 23, 2019, from https://en.wikipedia.org/w/index.php?title=Chromosome&oldid=907355235
Wikilectures. Prokaryotic Chromosomes (2017) In MediaWiki, Available at: https://www.wikilectures.eu/w/Prokaryotic_Chromosomes
Wikipedia contributors. (2019, May 15). DNA supercoil. In Wikipedia, The Free Encyclopedia. Retrieved 19:40, July 25, 2019, from https://en.wikipedia.org/w/index.php?title=DNA_supercoil&oldid=897160342
Wikipedia contributors. (2019, July 23). Histone. In Wikipedia, The Free Encyclopedia. Retrieved 16:19, July 26, 2019, from https://en.wikipedia.org/w/index.php?title=Histone&oldid=907472227
Wikipedia contributors. (2019, July 17). Nucleosome. In Wikipedia, The Free Encyclopedia. Retrieved 17:17, July 26, 2019, from https://en.wikipedia.org/w/index.php?title=Nucleosome&oldid=906654745
Wikipedia contributors. (2019, July 26). Human genome. In Wikipedia, The Free Encyclopedia. Retrieved 06:12, July 27, 2019, from https://en.wikipedia.org/w/index.php?title=Human_genome&oldid=908031878
Wikipedia contributors. (2019, July 19). Gene structure. In Wikipedia, The Free Encyclopedia. Retrieved 06:16, July 27, 2019, from https://en.wikipedia.org/w/index.php?title=Gene_structure&oldid=906938498